Content Sections
By Rob Verkerk PhD, founder, scientific and executive director, ANH-Intl
Alessandro had multiple myeloma. He had to decide whether or not to proceed with a second bone marrow transplant. He looked up the research in PubMed and found 4 trials that should have been able to answer his questions, but the data he really needed and that he knew the researchers would have known, wasn’t published. Alessandro isn’t a hypothetical person. We’re referring here to Alessandro Liberati PhD, the Italian healthcare researcher and clinical epidemiologist who founded and directed the Italian Cochrane Centre. Sadly, Alessandro passed away in 2012. But you’ll find an interview with him published by the Bulletin of the World Health Organization in 2010. He was one of a growing band of accomplished researchers with grave concerns about biases in research that result in the public being excessively and often unnecessarily medicated. His concerns brought him together with other researchers with similar concerns, including Dr Peter Gøtzsche, a co-founder of the Cochrane Collaboration who went on to expose organised crime and malfeasance within the medical research community, particularly where Big Pharma funding was involved.
The price paid for such whistleblowing is typically marginalisation by the crony-led ‘mainstream’ pharma-funded medical research community, although Alessandro never lived to see the full consequences of what he started.

Alessandro Liberati PhD. Source: BMJ
“Policy should be informed by evidence, not dictated by it: values and beliefs are an essential part of the decision making process. The key principle is transparency of decision-making” – associate professor Alessandro Liberati (2010) (now deceased), Medical School of the University of Modena and Reggio Emilia, Italy, and founder and director of the Italian Cochrane Centre.
Our own awakening about PubMed’s deficiencies
Alessandro Liberati, like Peter Gøtzsche, Peter Doshi, Tom Jefferson and others, including ourselves, have long been concerned about the concealment of research data that prevents independent analysis and therefore scientific interpretation. This means that clinical decisions and health policies are routinely made without full consideration of all relevant scientific data.
For most research scientists, the common starting point for evaluating the literature on biomedical issues is the online National Library of Medicine database, PubMed database, owned and operated by the US National Institutes for Health. The database claims to comprise “...more than 30 million citations for biomedical literature from MEDLINE, life science journals, and online books.”
PubMed is widely viewed as among the key databases used to conduct searches for relevant trials for inclusion in meta-analyses and systematic views. These kinds of studies are widely viewed as the pinnacle of the evidence-based medicine hierarchy and accordingly they are the most influential source of evidence in clinical decision-making.
But of course PubMed is used also by many others, including researchers, doctors, other clinicians, health policy makers, health journalists and even some citizens, including patients with serious diseases who are looking for answers and independent views of the science around their treatment options. Major public health decisions including those made by governments during this Covid-19 pandemic are based on science that is found in PubMed. It’s been the go-to engine for biomedical scientists for years – it is in many ways the Google of the biomedical world.
Unsurprisingly, independent non-profits engaged in the health space such as ourselves have also long been reliant on PubMed, and especially PubMed Central (PMC). But our view has been changing of late. Let me tell you why.
We’d been hearing about the imminent release of a new PubMed, something that was confirmed in a bulletin released by the US National Institutes for Health on 16 April. The incumbent version would become the legacy version on 18 May.
We also started to use the beta version of the new PubMed in early March. Four things immediately struck us during our research:
- The new PubMed lacked the dropdown menu of PMC or the legacy version which gives you access to the other databases such as PubMed Central (PMC), PubMed, MedGen, Books, etc.
- The number of articles found on the new PubMed following insertion of same search terms was often considerably less than when we searched the PMC database. This tendency was more noticeable when we searched for papers in controversial areas of medicine such as vaccines, or vitamin therapies for Covid-19 disease
- Sometimes key papers that we knew of – that we felt should have been found on the new PubMed (and were found on the PMC or Europe PMC) given the search terms used – were missing from the search result. We initially put this down to teething trouble. We're now getting more concerned about the algorithms behind the searches.
- Google and other search engines (e.g. DuckDuckGo) rapidly favoured the new PubMed engine in place of the legacy version that allows you to select PMC and other databases and we found we need to bookmark PMC or the legacy version in our internet browsers if we were to find them quickly.
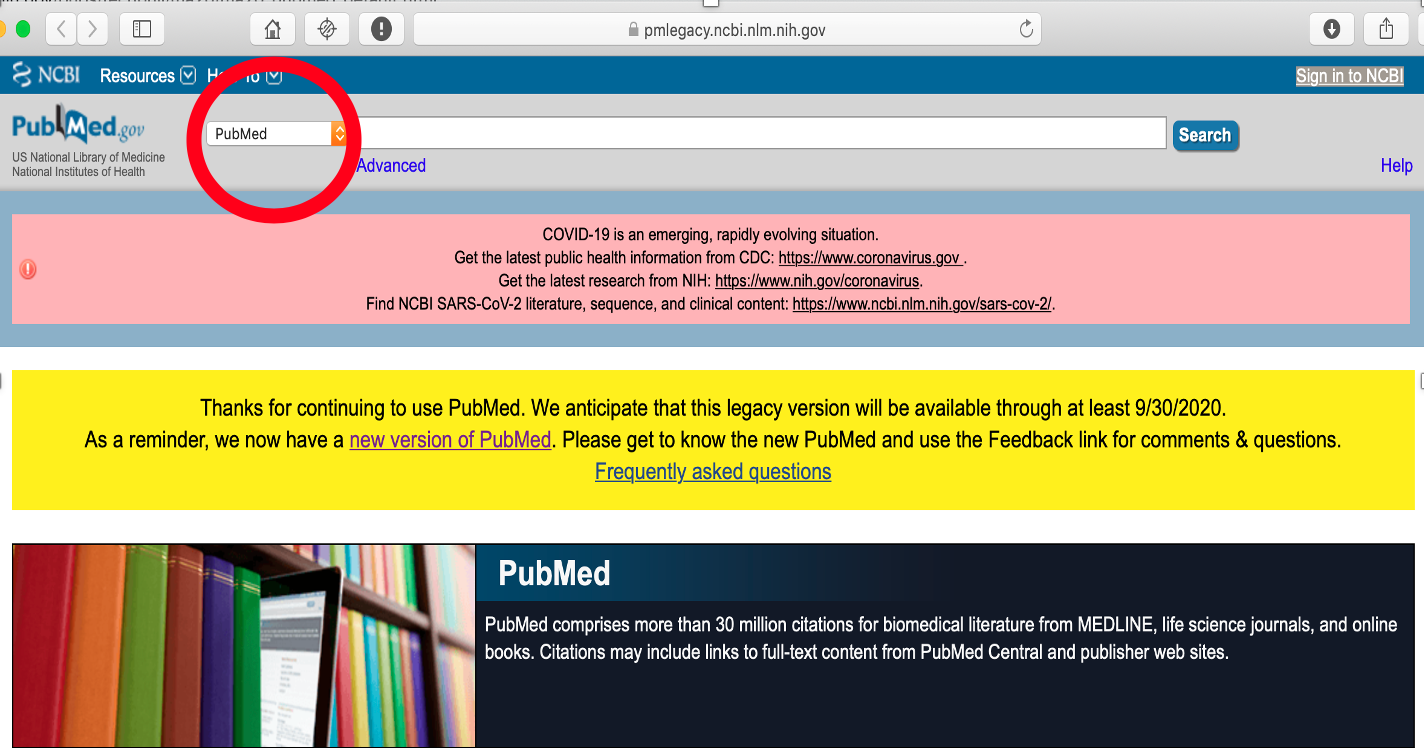
Front end of Legacy PubMed. Red circle shows dropdown menu that allows access to other databases, e.g. PMC.
Front end of New PubMed
So what happens if your database search engine doesn’t effectively capture all of the data and literature that has been published? What happens if you’re a biomedical researcher or an advisor to a government on health policy and you only see a part of the available information, all the while thinking your database is capturing everything?
What we found
Study 1: Searching controversial and uncertain scientific areas
During our weekly trawls of data around Covid for our covidzone.org science tracker that started in early March, we have found PubMed to be relatively useless. This is especially the case when we were searching for interactions between SARS-CoV-2 or Covid and natural agents such as vitamins. It was also very apparent when we searched for published literature around scientific transparency, particularly as it relates to vaccines. This latter issue was particularly relevant given our launch of a vaccine transparency campaign in late April.
So we conducted our own in-house investigation comparing the 4 key biomedical databases we commonly use or have used, namely the PubMed Central (PMC), Europe PMC, the new PubMed and the legacy PubMed version, across 6 different areas: heart disease, vaccines, natural medicines, micronutrients, genetically modified organisms (GMOs) and Covid. The summary results are shown in Figures 1 to 6 for each of these areas, respectively.
Hypothetically, if all 4 databases generated exactly the same number of hits, say, because they contained exactly the same references and the search engines were equally efficient at finding them, each database would generate 25% of the total number of hits for all 4 databases. Depicted as a pie chart, the pie would be divided into four equal quarters. On this basis, we’ve presented our results as pie charts so the relative distribution of references can be easily seen.
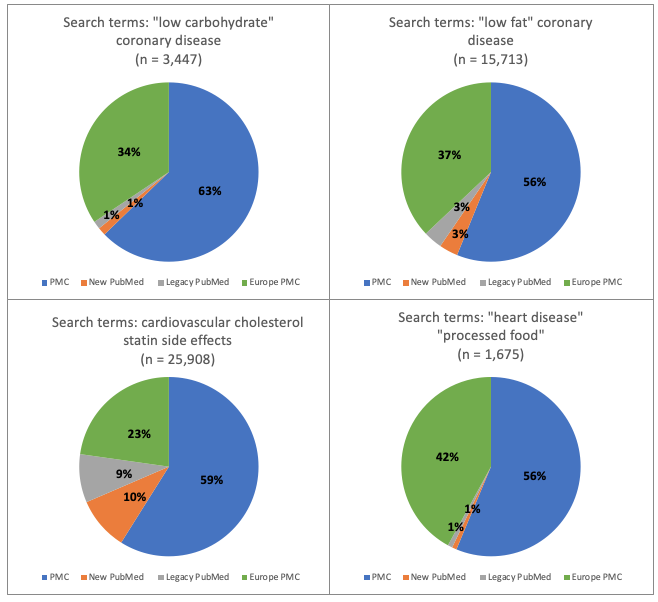
Figure 1. Research topic: heart disease. Percentage of total number of hits for each of 4 databases: new PubMed, legacy PubMed, PMC and Europe PMC using four different groups of search terms
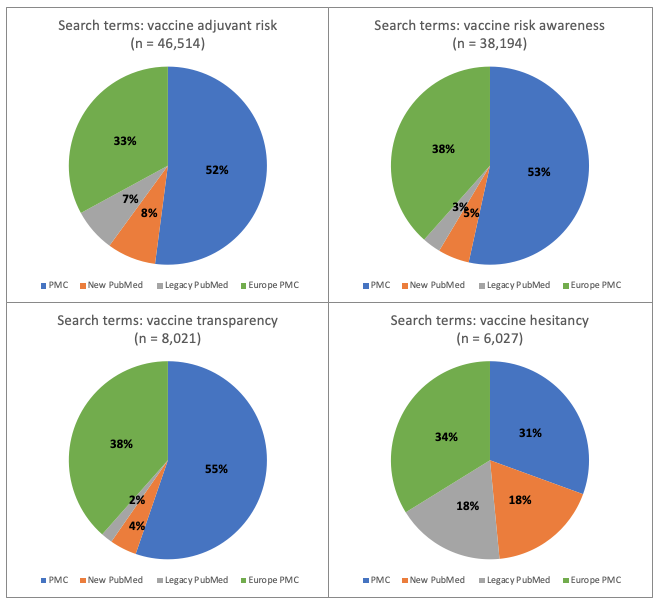
Figure 2. Research topic: vaccines. Percentage of total number of hits for each of 4 databases: new PubMed, legacy PubMed, PMC and Europe PMC using four different groups of search terms
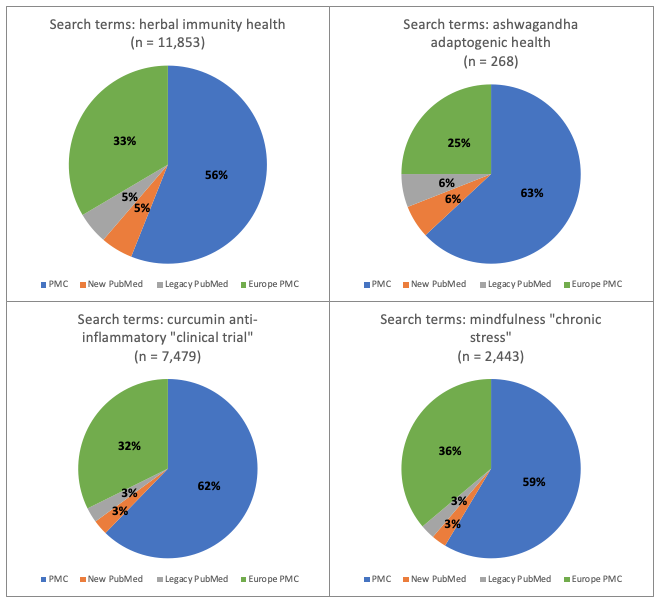
Figure 3. Research topic: natural medicines. Percentage of total number of hits for each of 4 databases: new PubMed, legacy PubMed, PMC and Europe PMC using four different groups of search terms
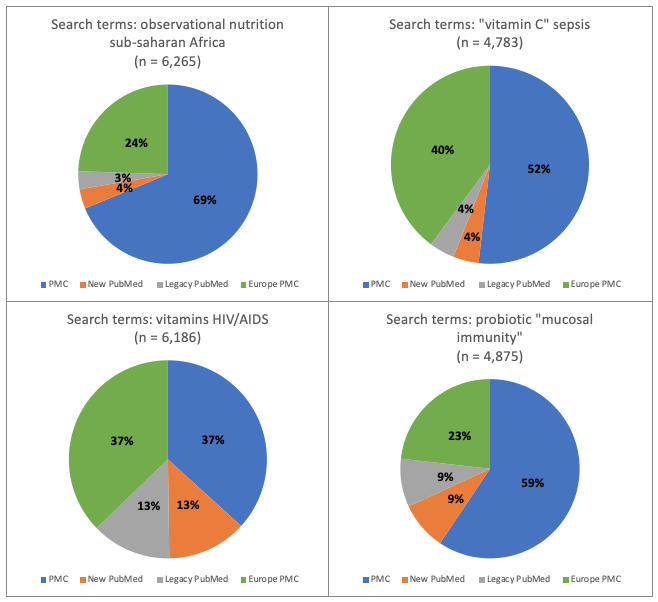
Figure 4. Research topic: micronutrients. Percentage of total number of hits for each of 4 databases: new PubMed, legacy PubMed, PMC and Europe PMC using four different groups of search terms
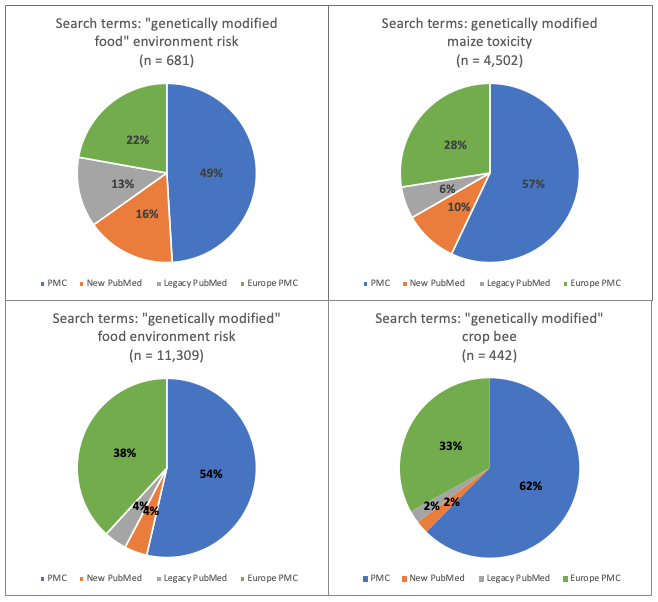
Figure 5. Research topic: genetically modified organisms (GMOs). Percentage of total number of hits for each of 4 databases: new PubMed, legacy PubMed, PMC and Europe PMC using four different groups of search terms
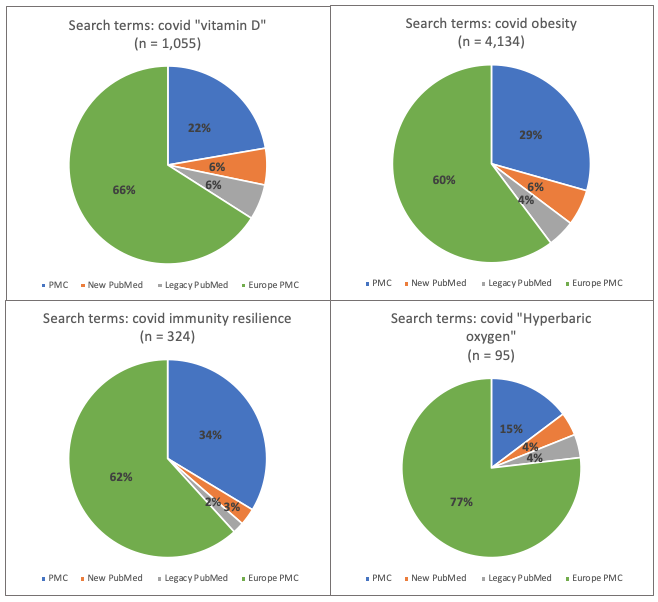
Figure 6. Research topic: Covid. Percentage of total number of hits for each of 4 databases: new PubMed, legacy PubMed, PMC and Europe PMC using four different groups of search terms
Our topline findings are as follows:
- PMC was the database that, with few exceptions, generated the largest number of hits for five out of the six areas, the exception being for Covid in which Europe PMC generated the largest number of hits
- Only one search in one area (vaccine hesitancy) showed an approximation to equivalency across the database
- With the exception of the 4 covid searches and the single vaccine hesitancy search, for all other searches the PMC database generated the greatest proportion (between 37% and 69%) of the total number of references across all databases
- Conversely, the PubMed databases each generated only between 1% and 18% of the total number of hits for all 4 databases, revealing they were the least efficient at gathering articles
- There were not big differences between the legacy and new versions of PubMed, although the new version generally generated fewer references
- We found incredibly significant, scientifically relevant papers, even some published in major high impact journals, that were missing on PubMed and were only found on the PMC or Europe PMC databases.
Study 2: Targeted searches of 4 databases
In order to understand better which references might be duplicated or missing between the 4 databases, we evaluated two different groups of search terms. The first (see summary results in Table 1) searched '"Jefferson T" transparency'. We chose this given that Tom Jefferson has been a major player in pushing for transparency of randomised trial data especially in relation to influenza and HPV vaccines. The second group of search terms (see summary results in Table 2) aimed to evaluate the science around what might be viewed as a more marginal (or controversial) area of covid science, namely the potential for using hyperbaric oxygen given that hypoxia (low blood oxygen levels) is common in patients seriously ill with Covid-19.
Table 1: Summary results from searches in 4 databases: PubMed (new version), PubMed (Legacy), PMC (PubMed Central) and Europe PMC with search terms: "Jefferson T" transparency
|
New PubMed |
Legacy PubMed |
PMC |
Europe PMC |
|
|
Total no of hits |
5 |
5 |
151 |
202 |
|
No of hits shared between all 4 databases |
1 |
|||
|
No of missing references in each database (found in alternate databases) |
10 |
11 |
4 |
5 |
|
Missing references |
Syst Rev; 9: 43 |
Recenti Prog Med; 108(1):7-10 |
Recenti Prog Med; 108(1):7-10 |
Recenti Prog Med; 108(1):7-10 |
1 Four papers were not found to be relevant as they omitted Jefferson T as author.
2 Six papers were found not to be relevant as they omitted Jefferson T as author.
Table 2: Summary results from searches in 4 databases: PubMed (new version), PubMed (Legacy), PMC (PubMed Central) and Europe PMC with search terms: covid "hyperbaric oxygen"
|
New PubMed |
Legacy PubMed |
PMC |
Europe PMC |
|
|
Total no of hits |
5 |
4 |
14 |
74 |
|
No of hits shared between all 4 databases |
2 |
|||
|
No of missing references in each database (found in alternate databases) 1 |
69 |
70 |
62 |
3 |
|
No of references in Europe PMC that are also found in 1 or more of the other databases |
14 |
- |
||
|
No of references found in Europe PMC that are missing from the other 3 databases |
- |
60 |
||
1 Reference list for most complete database (Europe PMC) available on request (please email [email protected]).
Frankly we were shocked by the inconsistency of results across the databases. We were able to draw the following conclusions:
- The searches confirmed that the two PMC databases yielded many more references than either of the PubMed databases
- In the first search, only 1 reference was common between all databases, in the second just 2
- There was a large number, proportionately, of missing references in the PubMed databases, compared with the PMC databases; these are shown for the first search (Table 1) as the number involved is manageable. In the second search, however, there were 60 references in the Europe PMC that did not appear in the other three databases.
Bottom line: if you use PubMed alone - you'll be blind to the science, at least in the areas we searched that are of particular interest to us and our supporters.
Conclusions
While many of us have glibly talked about searching PubMed for literature on all aspects of the biomedical sciences, we must now be aware that you should do this at your peril if you are genuinely interested in searching out all available peer reviewed literature. This might be especially the case when you carry out searches in areas that are viewed as marginal or a threat to mainstream interests.
We don't know where the funding came from to revise PubMed. But the removal of the dropdown menu is a disaster as it doesn't prompt the user to test hit rates using different databases. The funding for PubMed Central is, however, declared and we noted that at the top of the list of private and international partners is the omnipresent Bill and Melinda Gates Foundation. We simply don't know if this Foundation or any other interests were involved in the redevelopment of PubMed or the way in which PubMed is found by the internet search engines like Google. But given the recent revelations of Dr Philippe Douste-Blazy, a cardiologist and former French Health Minister, in an interview on French television who described disclosures made at a recent secret Chatham House rules meeting of experts. He upheld that the editor of The Lancet, Richard Horton, said that his journal and the New England Journal of Medicine were forced to publish a negative study on hydroxychloroquine following pressure put on them by the all-powerful pharmaceutical industry. He described the erosion of science as "criminal".
The process of now undertaking searches of PMC or Europe PMC - that we might have loosely referred to as a PubMed search (given PMC = PubMed Central) is now a convoluted process involving positively searching out PMC, rather the accessing it via the dropdown in the old (now legacy version).
Many people using the search engine may not have noticed what appears to be a sleight of hand. The National Center for Biotechnology Information (NCBI) will undoubtedly argue that it made it abundantly clear what was happening in its 16 April bulletin.
Whatever the reasons, we've demonstrated that PubMed is next to useless as a database and resource for the kind of subject areas central to our mission and vision.
We're not aware of any other individual or organisation that's sounded this warning. So please, share this widely, especially among doctors, clinicians or practitioners who may still be likely to use PubMed as their key data source.
IMPORTANT NOTE
Please share this article widely. We are currently facing serious censorship from social media platforms so our ability to communicate is heavily dependent on what you, the reader, decides to do, or not do, after reading the article. As an independent non-profit focused on natural and sustainable health, our survival as an organisation is entirely dependent on donations. We greatly welcome donations, however big or small, via our secure online portal. Ethical companies involved in the natural sector can join our Vanguard Club and derive unique benefits. Thank you.
>>> Back to Alliance for Natural Health Intl homepage








Comments
your voice counts
18 June 2020 at 8:59 am
This is horribly disturbing! I have posted a link to this article on both the GCRN and ARCH F.B. pages.
18 June 2020 at 12:29 pm
Thank you for sharing Richard, we appreciate your support.
Warm regards
Melissa
18 June 2020 at 9:01 am
Dear ANH, please sue or seek prosecution of those responsible. Thank you.
18 June 2020 at 10:53 am
Control of the narrative is also the control of the definitions from which diagnostic judgements are based.
Like banker's toxic debts, we have a scientific literature in which perhaps half or more is not only worthless but used as a foundational basis from which further funding, research, and policy development operates. Like banker's debt, the mind that fears loss under transparency of account, seeks to reset the system so as to set the foundation for its private gain or 'salvation' from fear, pain and loss.
Whatever we search for, we search with, and transparency to our own account is a basis for operating on just and true premises rather than self-deceiving obfuscations and cognitive dissociations that render stuffed presentations in hollow parody of life.
18 June 2020 at 11:10 am
Clearly this is deeply worrying, another demonstration of how the pharmaceutical industry (and it 'philanthropic' allies) are subverting medical science.
I had thought that by using 'DuckDuckGo' I was using a search engine less likely to be involved in this.
Do you have any information about search engines that are more likely to 'find' the kind of evidence you and I regularly search for?
18 June 2020 at 6:16 pm
A brilliant analysis, which mirrors exactly the findings by Sayer Ji, Ty & Charlene Bollinger et al in the United States. Are we still expected to believe that there is no conspiracy and that nobody is pulling the strings?
18 June 2020 at 7:57 pm
Thanks for the comment Werner. We weren't aware that Sayer Ji or Ty and Charlene B had commented on the problems with PubMed. Any links to their info would be helpful, thanks. Best, Rob
19 June 2020 at 12:16 pm
As always, well researched, well presented and such thoughtful, highly intelligent comments. Thank you all at ANH I'nal! I also receive the Sayer Ji (have for years) & C & T Bollinger emails, but always the standard is so high here at ANH I'nal.
I too was surprised re "DuckDuckGo", oh well, another email to be written - so many for some years now, about lack of standards, corruption and I now ask outright or infer that they are on Bill Gates, Soros, Oprah Winfrey, Rockefeller and fellow travellers payrolls - of course, I realise that they are simply through their FOUNDATIONs being Charitable and thinking only of their (fellow) being.
I keep telling people that the Mental Hospitals as they were called when I was little, and no longer exist, is one reason that we see so many Sociopaths loose in Society, but I never thought it as going to be an epidemic in itself.
Also received Robert F Kennedy Jnr "Children's Health Defense.org" email this am, there is one Video/Article in particular that is REQUIRED reading today. It always, like ANH full of "spot on" information, but todays video is unbelieavable as to what Scientists can be involved in.
RFK, Jr. Discusses "Aborted Foetal Tissue and Vaccines" and so much more related scientific information, with Dr. Theresa Deisher (Scientist) who established and runs this not for profit organisation: www.soundchoice.org
This Scientist is so knowledgeable about Vaccines, history of and so much more.
Even RFK Jnr was shocked more than once during this Interview. bear in mind that RFK Jnr is a Lawyer and has proved his commitment to human beings already, and now is concerned about Autism and Vaccines of course.
P.S. Brain Steere, you have shocked me "stuffed" presentations. You always have such a command of the English language too. It sounds as though you too, are totally "fed up" with all this pathetic, destructive behaviours of the few onto the many.
19 June 2020 at 12:41 pm
Addendum: Has a Schematic Diagram ever been done re Gates "Donations " to Universities, Research Laboratories, Companies he has bought, others that he makes "Donations" to and so on. A picture tells a thousand words and this would be so useful to distribute to get the message to people who doubt just how much this criminal has been planning and doing for many so many years, and one would need for Indians, Africans, Pakistanis and so many other people/Nationalities to contribute towards this Schematic Diagram?
I do know that University of Queensland is a benefactor and caught up in the web re getting a Vaccine for Moderna out into the Market place or one of the other 8 Vaccine factories that Gates owns, or maybe that Oracle of the media, who converted to Americanism, even down to the Texan wife, Rupert Murdoch who owns the largest Vaccine Company in Australia if my source is correct may contribute to all things Australian.
But I agree it would take a Battalion of people to research and list all and maybe even then not find holdings he has or beneficent donations given.
21 July 2020 at 10:52 pm
Deirdre Ryan, I agree completely. There's a part of my brain that has been designing a diagram of the web of money spilling from his foundation for the past few months now. My main concern is how messy it would be with lines criss-crossing back and forth. It needs someone with a bit of artistic creativity to design a map of this corruption and sadly my creativity lies in other areas! Great idea. Maybe someone can help?
07 November 2022 at 7:22 pm
It's all just like George Orwell's 1984...to control the narrative. I've only just subscribed and really appreciate your exposure on this "erosion of science"...we all need to think critically. I will pass this on. Keep up the honest work.
Your voice counts
We welcome your comments and are very interested in your point of view, but we ask that you keep them relevant to the article, that they be civil and without commercial links. All comments are moderated prior to being published. We reserve the right to edit or not publish comments that we consider abusive or offensive.
There is extra content here from a third party provider. You will be unable to see this content unless you agree to allow Content Cookies. Cookie Preferences